European Training Network coordinated at CRG in Barcelona
OPATHY is an innovative translational research training network coordinated by the CRG that will explore the potential of next-generation high-throughput technologies to study the interactions of yeasts that cause human diseases.Identification of pathogenic yeasts
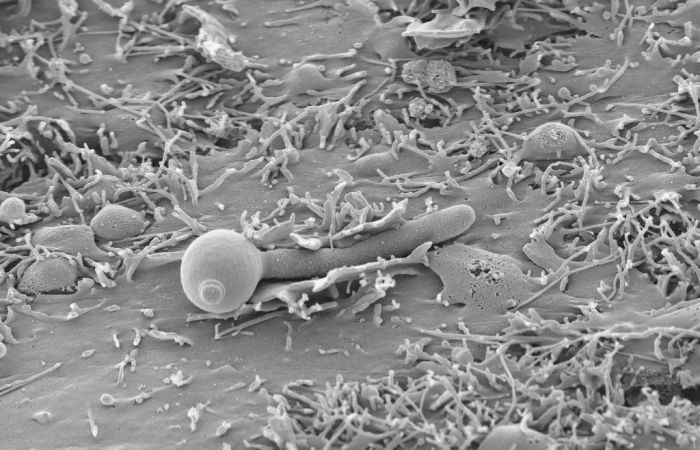
Candida tropicalis has been identified as the most prevalent pathogenic yeast species of the Candida-non-albicans group. Historically, Candida albicans has been the major species responsible for causing candidiasis.Third annual meeting Aug/ Sep 2018 at Biotechvana, Spain
The 3rd OPATHY annual meeting took place from 26th of August to 1st of September 2018 at the University of Valencia Science Park, Spain and had a focus on applied bioinformatics and post-genomics techniques for clinical omics.Training and Networking Activities
The designed OPATHY training program will provide the fellows with the necessary expertise to improve their research skills and enhance their career prospects.

OPATHY
From Omics to Patient: Improving Diagnostics of Pathogenic Yeasts